Many analysis tools deal with networks in the form of an adjacency matrix, and exposing the aMatReader API to automation users enables scripts to transfer those networks directly into Cytoscape with little effort.Īdjacency Cytoscape Interoperability Microservice REST Reproducibility Workflow. We also exposed CyREST endpoints to allow researchers to import network matrix data directly into Cytoscape from their language of choice. The input file must have the '.mat' extension to avoid conflict with other readers and the format of the file should have the names of the nodes in the top for and as the first field in each row thereafter. To accelerate the import process, aMatReader attempts to predict matrix import parameters by analyzing the first two lines of the file. aMatReader is a Cytoscape 3 app that augments the Cytoscape network readers by adding the ability to read in an adjacency matrix. from the tutorial files provided in the PathBLAST plugin of software Cytoscape 1.1 was. Converts an adjacency matrix to Cytoscape's sparse matrix format Usage convert2cytoscape(A) Arguments. Our goal was to import the diverse adjacency matrix formats produced by existing scripts and libraries written in R, MATLAB, and Python, and facilitate importing that data into Cytoscape. The aMatReader app enables users to import one or multiple adjacency matrix files into Cytoscape, where each file represents an edge attribute in a network. Note that the elements of adjacency matrices may be symmetric or. 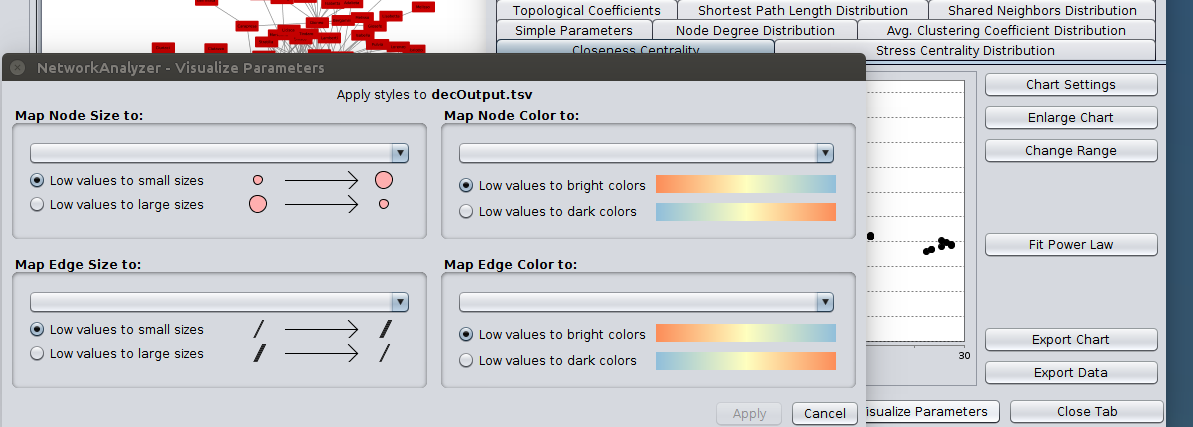
The aMatReader app enables users to import one or multiple adjacency matrix files into Cytoscape, where each file represents an edge attribute in a network. How to make a network diagram using Cytoscape Dear experts, I want to make a network diagram for metabolites and genes based on PCC (Pearson Correlation Coefficient) value. Adjacency matrices are useful for storing pairwise interaction data, such as correlations between gene pairs in a pathway or similarities between genes and conditions.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
